Team Discovers Genes That Turn Horses Into Racehorses
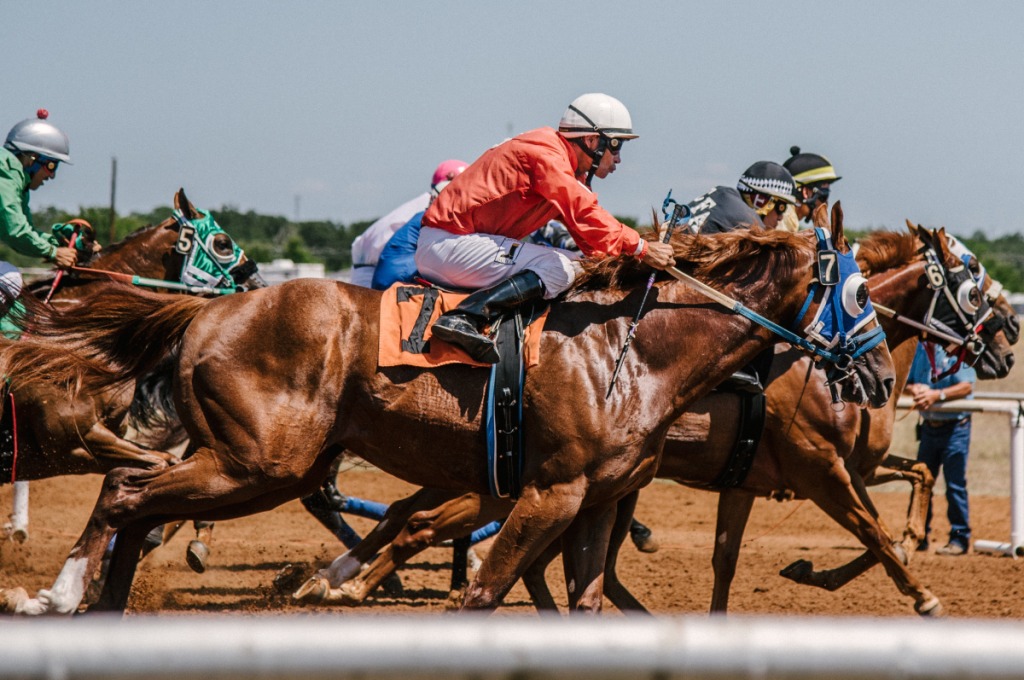


Racehorses really are a breed apart, according to new research which has identified the genes that make them into good racers.
They fuel sprinting ability, stamina, muscle, metabolism and brainpower – and differ from variants in their less speedy peers.
It confirms the world’s greatest names like Red Rum, Arkle, Frankel and Desert Orchid are all distantly related.
An international team compared the complete DNA of Thoroughbred, Arabian and Mongolian racehorses to those bred for other sports and leisure activities. The study is published in the journal Communications Biology.
Lead author Professor Emmeline Hill, of University College Dublin (UCD), said: “The very large number of horse breeds developed over the last hundreds of years all over the world have been carefully shaped by selective breeding for different traits desired by breeders.
“This has led to tall horses, small horses, powerful draft horses, useful riding horses, and fast racing horses.”
Making breeding less of a lottery is the “holy grail” for owners and trainers. Knowing what genes help horses sprint longer distances could lead to higher stud fees.
Hill said: “We have discovered a set of genes common to racing horses, but not all horses within a racing breed have the advantageous gene version.
“So these findings will be useful to identify the most suitable individuals within a breed for racing or for breeding.”
The mutated versions were found to be common to all racing breeds – and clearly different to equivalents in non-racing breeds.
Hill, who is also chief science officer at Dublin-based equine sports company Plusvital, said: “Since the discovery of the ‘Speed Gene’ in 2009, we have generated genetic data for thousands of Thoroughbreds and horses from other breeds

“This is the first time this set of genes has been linked to the success of racing breeds. Two of the genes were previously identified for performance in Thoroughbreds and Arabians.
“But the approach we took was to ask what genes were common to all racing breeds and different from non-racing breeds.”
Thoroughbred horses are finely-tuned “athletes” with a high aerobic capacity relative to their skeletal muscle mass – attributed to centuries of genetic selection for speed and stamina.
Non-genetic factors such as variations in training schedule can also influence how racehorse distance aptitudes and preferences develop.
Co0author Professor David MacHugh, also from UCD, said: “Although racing is a multifactorial trait, with management and training having a considerable influence on the success of a racehorse, this study provides good evidence for major-effect genes shaping the racing trait in horse populations.”
The researchers collected hair samples from 100 horses owned by the champion Ajnai Sharga Horse Racing Team at their breeding farm in Khentii province, Mongolia, the birthplace of Ghengis Khan.
Using the DNA from these Mongolian racehorses, along with Thoroughbred and racing Arabian horses, the scientists compared the genomes with 21 other non-racing breeds.
They included the Clydesdale, Connemara pony, Hanoverian, Morgan, Norwegian Fjord, Paint, Shetland and Shire. The analysis identified seven essential genes for racing.
Among the most important was NTM, which functions in brain development and influences learning and memory.
It was selected during the horse domestication process. In Thoroughbreds, it influences whether a horse ever races.
MacHugh said: “This finding suggests equine neurological systems perturbed by natural and artificial selection associated with domestication may overlap with adaptive traits that are required for racing.”
Inside knowledge of what makes the fittest horses tick is potentially priceless in the multi-billion dollar industry.
The initial breakthrough by the same team more than a decade ago led to speculation about the creation of superstar racehorses – in the lab.
Many old school breeders were averse to science, preferring to concentrate on look and lineage. But younger rivals are embracing the challenge.
First author Dr. Haige Han, also from UCD, said: “Testing these variants in new sets of hundreds of horses from racing and non-racing breeds identified seven essential genes for racing.
“These genes have roles in muscle, metabolism, and neurobiological functions, and are central to racing ability among horse breeds.”
The researchers used data from Thoroughbred horses to investigate if the genes they had identified were involved in the muscle response to exercise and training.
Hill added: “By integrating these two different data sets we fine-tuned the list of racing genes to those that were most biologically relevant to racing.
“One of these genes was MYLK2 which is required for muscle contraction. In humans, MYLK2 is associated with exercise-induced muscle damage.”
Produced in association with SWNS Talker.
The Western Journal has not reviewed this story prior to publication. Therefore, it may not meet our normal editorial standards. It is provided to our readers as a service from The Western Journal.
Advertise with The Western Journal and reach millions of highly engaged readers, while supporting our work. Advertise Today.
Truth and Accuracy
We are committed to truth and accuracy in all of our journalism. Read our editorial standards.